Secondary (Scatter) Radiation and Undercut Control
Secondary (Scatter) Radiation
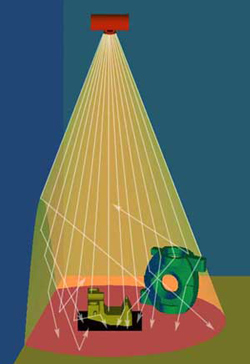 Secondary or scatter radiation must often be taken into consideration when producing a radiograph. The scattered photons create a loss of contrast and definition. Often secondary radiation is thought of as radiation striking the detector reflected from an object in the immediate area, such as a wall, or from the table or floor where the part is resting. Side scatter originates from walls, or objects on the source side of the detector. Control of side scatter can be achieved by moving objects in the room away from the detector, moving the x-ray tube to the center of the vault, or placing a collimator at the exit port, thus reducing the diverging radiation surrounding the central beam.
Secondary or scatter radiation must often be taken into consideration when producing a radiograph. The scattered photons create a loss of contrast and definition. Often secondary radiation is thought of as radiation striking the detector reflected from an object in the immediate area, such as a wall, or from the table or floor where the part is resting. Side scatter originates from walls, or objects on the source side of the detector. Control of side scatter can be achieved by moving objects in the room away from the detector, moving the x-ray tube to the center of the vault, or placing a collimator at the exit port, thus reducing the diverging radiation surrounding the central beam.
Backscattered x-rays can be significant when using film as a detector. They come from objects behind the film. They are NOT important in digital detectors. Industry codes and standards often require that a lead letter "B" be placed on the back of the film cassette to verify the control of backscatter. If the letter "B" shows as a "ghost" image on the film, a significant amount of backscatter radiation is reaching the film. The image of the "B" is often quite non-distinct because the backscatter source is diffuse. The control of backscatter radiation is often achieved by backing the film in the cassette with a sheet of lead that is at least 0.010 inch thick. It is a common practice in industry to place a 0.005" lead screen in front and a 0.010" screen behind the film.
Undercut
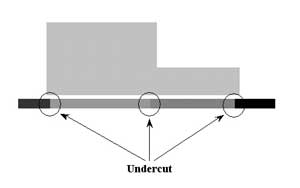 Another condition that must often be controlled when producing a radiograph is called undercut. Parts with holes, hollow areas, or abrupt thickness changes are likely to suffer from undercut if controls are not put in place. Undercut appears as a darkening of the film radiograph in the area of the thickness transition. It appears as a brighter ghosting near the edges in a digital image. This results in a loss of resolution or blurring at the transition area. Undercut occurs due to scattering within the detector. At the edges of a part, or areas where the part transitions from thick to thin, the intensity of the radiation reaching the detector is much greater than in the thicker areas of the part. The high level of radiation intensity reaching the detector results in a high level of scattering within the detector. Using film, it should also be noted that the faster the film speed, the more undercut that is likely to occur. Scattering from within the walls of the part also contributes to undercut, but research has shown that scattering within the detector is the primary cause. Shutters or masks can be used to reduce undercut. Sheets of lead cut to fill holes or surround the part and metallic shot and liquid absorbers are often used as masks.
Another condition that must often be controlled when producing a radiograph is called undercut. Parts with holes, hollow areas, or abrupt thickness changes are likely to suffer from undercut if controls are not put in place. Undercut appears as a darkening of the film radiograph in the area of the thickness transition. It appears as a brighter ghosting near the edges in a digital image. This results in a loss of resolution or blurring at the transition area. Undercut occurs due to scattering within the detector. At the edges of a part, or areas where the part transitions from thick to thin, the intensity of the radiation reaching the detector is much greater than in the thicker areas of the part. The high level of radiation intensity reaching the detector results in a high level of scattering within the detector. Using film, it should also be noted that the faster the film speed, the more undercut that is likely to occur. Scattering from within the walls of the part also contributes to undercut, but research has shown that scattering within the detector is the primary cause. Shutters or masks can be used to reduce undercut. Sheets of lead cut to fill holes or surround the part and metallic shot and liquid absorbers are often used as masks.